You don’t have to be a scientist to do science.
By simply running a free program, you can help advance research in medicine, clean energy, and materials science.
|
|
How does it work?
By running Rosetta@home on your computer when you're not using it you will speed up and extend our efforts to design new proteins and to predict their 3-dimensional shapes. Proteins are the molecular machines and building blocks of life. You can read more about protein folding and design here.
Follow us on Twitter: @rosettaathome
Rosetta@home is not for profit.
Join Ralph@home to help improve this project.
User of the Day
Predictor of the day
Predictor of the day: Congratulations to 2buiy5RiX3bN8JBr2yuiTCdV5yLJ (Team Gridcoin) for predicting the lowest energy structure for workunit rb_02_26_22117....
1 Mar 2022, 0:00:00 UTC
· Discuss
... more
Predictor of the day is available as an RSS feed
News
COVID-19 vaccine with IPD nanoparticles wins full approval abroad
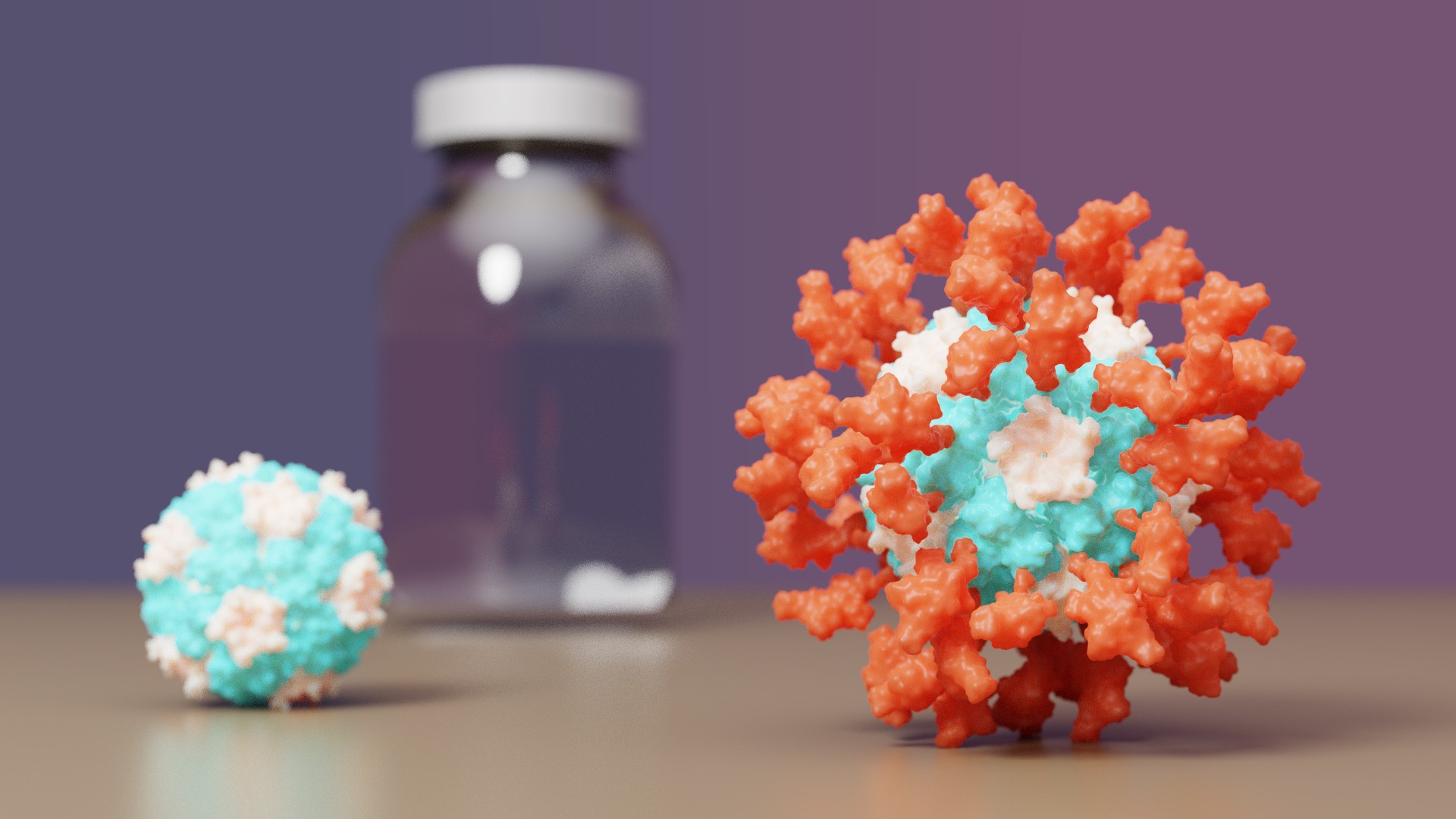
The IPD is excited to announce it's first designed protein medicine with full approval abroad.
Congrats and thank you to all Rosetta@home contributors! The computing you have provided has greatly aided in de novo protein design challenges such as vaccine development leading to breakthroughs like this.
For more information you can visit the IPD vaccine news post.
A video is also available here.
From the IPD news site:
• Clinical testing found the vaccine outperforms Oxford/AstraZeneca’s
• The protein-based vaccine, now called SKYCovione, does not require deep freezing
• University of Washington to waive royalty fees for the duration of the pandemic
• South Korea to purchase 10 million doses for domestic use
A vaccine for COVID-19 developed at the University of Washington School of Medicine has been approved by the Korean Ministry of Food and Drug Safety for use in individuals 18 years of age and older. The vaccine, now known as SKYCovione, was found to be more effective than the Oxford/AstraZeneca vaccine sold under the brand names Covishield and Vaxzevria.
SK bioscience, the company leading the SKYCovione’s clinical development abroad, is now seeking approval for its use in the United Kingdom and beyond. If approved by the World Health Organization, the vaccine will be made available through COVAX, an international effort to equitably distribute COVID-19 vaccines around the world. In addition, the South Korean government has agreed to purchase 10 million doses for domestic use.
The Seattle scientists behind the new vaccine sought to create a ‘second-generation’ COVID-19 vaccine that is safe, effective at low doses, simple to manufacture, and stable without deep freezing. These attributes could enable vaccination at a global scale by reaching people in areas where medical, transportation, and storage resources are limited.
“We know more than two billion people worldwide have not received a single dose of vaccine,” said David Veesler, associate professor of biochemistry at UW School of Medicine and co-developer of the vaccine. “If our vaccine is distributed through COVAX, it will allow it to reach people who need access.”
The University of Washington is licensing the vaccine technology royalty-free for the duration of the pandemic.
Congrats and thank you again to all R@h contributors!
6 Jul 2022, 18:27:59 UTC
· Discuss
Thank you!
We'd like to thank everyone who has contributed and continues to contribute to this project, and would like to remind everyone that there may be periods of down time while we are preparing for future large scale batches of jobs and analyzing results. We greatly appreciate your contributions which are vitally important to our ongoing research.
Thank you!
5 Nov 2020, 20:16:29 UTC
· Discuss
Coronavirus update from David Baker. Thank you all for your contributions!
Here is a short video of David Baker describing some exciting results from de novo designs targeting SARS-Cov-2.
Thank you all for your contributions to this research! Although R@h was not directly used for the work described in the publication (link provided below), R@h was used for designing relevant scaffolds. Additionally, there are currently many similar designs that bind SARS-Cov-2 and related targets that were engineered using R@h.
https://www.youtube.com/embed/ODEIN5V3yLg
More information is available from the publication, De novo design of picomolar SARS-CoV-2 mini protein inhibitors.
21 Sep 2020, 23:16:33 UTC
· Discuss
Switch to using SSL (Secure Socket Layer)
We updated our project to use SSL. The project URL has thus been changed to https://boinc.bakerlab.org/rosetta. You can reattach the project using this updated URL at your convenience. Please post any issues regarding this update in the discussion thread.
1 May 2020, 21:25:30 UTC
· Discuss
Help in the fight against COVID-19!
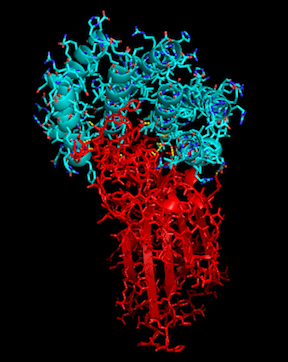
With the recent COVID-19 outbreak, R@h has been used to predict the structure of proteins important to the disease as well as to produce new, stable mini-proteins to be used as potential therapeutics and diagnostics, like the one displayed above which is bound to part of the SARS-CoV-2 spike protein.
To help our research, we are happy to announce a new application update, and thanks to the help from the Arm development community, including Rex St. John, Dmitry Moskalchuk, David Tischler, Lloyd Watts, and Sahaj Sarup, we are excited to also include the Linux-ARM platform. With this update we will continue to make protein binders to SARS-CoV-2 and related targets using the latest Rosetta source.
Thank you R@h volunteers for your continued support to this project. Your CPU hours are used not only to accurately model the structures of important proteins, but to design new ones as well. Let's band together and fight COVID-19!
More details will be available in the Discussion of this news post.
3 Apr 2020, 3:45:24 UTC
· Discuss
... more
News is available as an RSS feed

©2024 University of Washington
https://www.bakerlab.org